Research Project 2: Summary
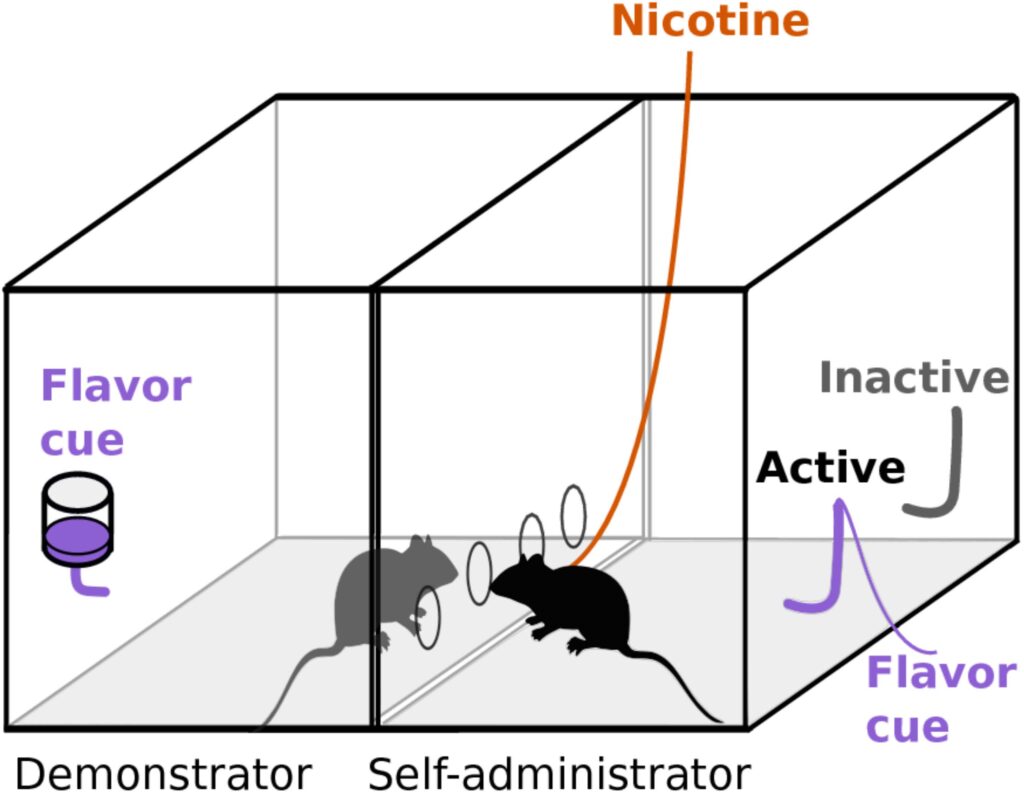 Tobacco use is the single most preventable cause of disease, disability, and death in the United States. Clinical studies have demonstrated that heritability is ~50% for various smoking behaviors. However, most of the genetic factors determining smoking behavior remain unknown. Genetic studies using rodent models have greatly improved our understanding of the cause of human genetic diseases and discovered many targets for therapeutic drugs. We have established a rat model in which social cue is an important determinant of nicotine intake. This model is particularly pertinent to smoking because it incorporates the critical effect of social environment and the aversive effect of nicotine. Both of these effects are well recognized in clinical investigations, but are largely ignored in preclinical studies. The heterogenous stock (HS) rats are particularly suitable for fine genetic mapping studies. Therefore, the primary goal of Research Project 2 is to phenotype 3,200 HS rats on socially-acquired nicotine self-administration. We use adolescent HS rats bred locally to capture the role of this critical brain development stage on smoking initiation. We also test these HS rats on contextual-cue induced reinstatement of drug seeking. Data on several social and emotional traits is obtained before self-administration starts. Multiple regression will be used to analyze the relationship between these traits with nicotine intake and reinstatement. DNA samples from each rat will be sent to the Core C for genotyping, as well as quantitative behavioral measures to perform a genome-wide association study (GWAS) on nicotine intake and reinstatement of nicotine seeking. Sex difference will be examined. We also precisely collected tissue from 88 HS rat brains. Five brain regions that are critical for the behaviors investigated by Research Projects 1-3 were targeted: nucleus accumbens core, lateral habenula, orbitofrontal cortex, infralimbic cortex, and prelimbic cortex. Core C will analyze gene expression data and integrate the results with the behavioral and genomic findings.
Tobacco use is the single most preventable cause of disease, disability, and death in the United States. Clinical studies have demonstrated that heritability is ~50% for various smoking behaviors. However, most of the genetic factors determining smoking behavior remain unknown. Genetic studies using rodent models have greatly improved our understanding of the cause of human genetic diseases and discovered many targets for therapeutic drugs. We have established a rat model in which social cue is an important determinant of nicotine intake. This model is particularly pertinent to smoking because it incorporates the critical effect of social environment and the aversive effect of nicotine. Both of these effects are well recognized in clinical investigations, but are largely ignored in preclinical studies. The heterogenous stock (HS) rats are particularly suitable for fine genetic mapping studies. Therefore, the primary goal of Research Project 2 is to phenotype 3,200 HS rats on socially-acquired nicotine self-administration. We use adolescent HS rats bred locally to capture the role of this critical brain development stage on smoking initiation. We also test these HS rats on contextual-cue induced reinstatement of drug seeking. Data on several social and emotional traits is obtained before self-administration starts. Multiple regression will be used to analyze the relationship between these traits with nicotine intake and reinstatement. DNA samples from each rat will be sent to the Core C for genotyping, as well as quantitative behavioral measures to perform a genome-wide association study (GWAS) on nicotine intake and reinstatement of nicotine seeking. Sex difference will be examined. We also precisely collected tissue from 88 HS rat brains. Five brain regions that are critical for the behaviors investigated by Research Projects 1-3 were targeted: nucleus accumbens core, lateral habenula, orbitofrontal cortex, infralimbic cortex, and prelimbic cortex. Core C will analyze gene expression data and integrate the results with the behavioral and genomic findings.
RatsPub: a webservice aided by deep learning to mine PubMed for addiction-related genes
Updated: October 2020
