We share data in these ways: Internet Rat Server | Datasets | RatGTEx Portal | SRA archive | UC San Diego Library Digital Collections | GitHub | Biobanks | GeneNetwork2
Internet Rat Server (IRS)

The Internet Rat Server (IRS) is a cloud-hosted portal providing the Center’s investigators with easy, password-protected access to the Center’s HS rats database, including an archive of reports and data tools.
The Center maintains this database containing all phenotypical data collected by the HS rats research projects and ancillary collaborators, as well as RNAseq and genotype data.
Data requests should be addressed to the Center PI, Abraham Palmer or the Scientific Coordinator, Oksana Polesskaya.
Datasets

Dataset lists from publications and projects related to the Center are available here on ratgenes.org, and are designed to comply with FAIR (Findable, Accessible, Interoperable, Reusable) data principles.
(Note: previously available on the C-GORD portal, which was active 2021-2025.)
RatGTEx Portal

RatGTEx Portal provides interactive visualizations and data downloads for queried genes, variants, and rat tissues. It will specify a standard processing pipeline and data formats to facilitate comparisons across datasets.
(See more on our eQTL mapping page.)
Raw sequence reads

Sequencing data for HS rats: BioProjects are deposited to the NIH Sequence Read Archive (SRA).
Low-coverage whole-genome sequencing:
- N/NIH heterogeneous stock (HS) outbred rats, low-coverage whole genome sequencing (PRJNA1022514)
Deep-coverage whole-genome sequencing (using short-reads lllumina method):
- Whole-genome sequence data for 88 outbred heterogeneous stock rats (PRJNA1076141)
- Whole-genome sequence data for 8 heterogeneous stock rat founders (PRJNA1048943)
UC San Diego Library Digital Collections

PalmerLab GitHub

PalmerLab GitHub contains releases of genotyping bioinformatics pipelines and other related projects. The content is updated as the lab develops new pipelines.
Addiction Biobanks
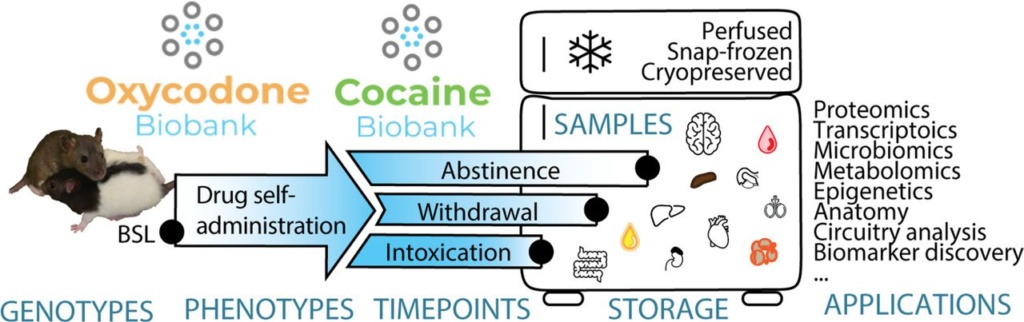
We are working with UC San Diego researchers Dr. Olivier George and Dr. Giordano de Guglielmo to provide genotyping for the HS rats that underwent cocaine, oxycodone, or alcohol exposure protocols.
The tissues from these animals were collected and preserved in tissue banks and are available for researchers around the world to use in addiction studies.
Request forms are available on the Addiction Biobank website:
- cocaine biobank
- oxycodone biobank
- alcohol biobank (more info at De Guglielmo Lab)
The Cocaine and Oxycodone Biobanks, Two Repositories from Genetically Diverse and Behaviorally Characterized Rats for the Study of Addiction. eNeuro 19 April 2021; 8 (3) ENEURO.0033-21.2021.
GeneNetwork

We are working with the GeneNetwork 2 team at the University of Tennessee Health Science Center to make our data available through this resource. GeneNetwork2 provides access to linked datasets and tools to study complex networks of genes, molecules, and higher-order gene function and phenotypes.
Researchers interested in obtaining additional data from this center should contact the Center PI, Abraham Palmer.
Updated on:
